When a new COVID-19 variant appears, such as the most recent Omicron variant of concern, scientists want to learn as much about it, in as short a time as possible. More than just understanding the genetic changes to the variant, scientists want to anticipate the potential impacts, such as whether it will spread more easily, or cause more severe disease.
Dr. Daniel Beniac, a scientist at the Public Health Agency of Canada’s National Microbiology Laboratory (NML), creates three-dimensional (3D) computer models of COVID-19 variants to predict if certain mutations lead to a more infectious virus.
Some viruses, including the virus that causes COVID-19, have spike proteins on their surface (crown-like structures on the surface of the virus) which allow it to bind to a specific protein on the surface of a cell and subsequently infect the cell. This computer modelling system generates 3D models to virtually assess if certain mutations in the spike protein of the COVID-19 virus create improved binding. If mutations make it easier for the virus to bind to a cell, it could mean that the virus can infect more easily. While this method does not look at the impact of mutations elsewhere in the virus, it does help to calculate how the mutation behaves.
Electron microscopes provide starting point
An electron microscope is used to generate high-resolution images. Electron microscopes work by accelerating an electron beam and firing this through the specimen. To conduct this work, spike and cell receptor proteins are fabricated in the lab, purified, imaged in the electron microscope and analyzed in a computer. This results in the production of a 3D structure of the spike binding protein, which is the starting point for creating a 3D model. This structure can then serve as a platform for generating 3D models of any new variant as they arise. To do this a computer inserts the mutations found in a variant, and then they are analyzed to measure the effect of the variant.
During the development phase, Dr. Beniac often worked long hours doing manual data entry processing hundreds to thousands of spike models for each variant to test receptor binding. Eventually, the manual work was replaced with an automated process that runs on a powerful computer. This large data processing ability made it possible to achieve the goal of a 1 to 2-day turnaround time for results that once took days or weeks.
“One of the real bonuses here in the NML is that we have the virology and protein biology part of the lab that lets us work with these pathogens and proteins. Then we can use electron microscopy and computational analysis to study the structural biology end of it,” said Dr. Beniac. “We’ve got the best of both worlds at the NML.”
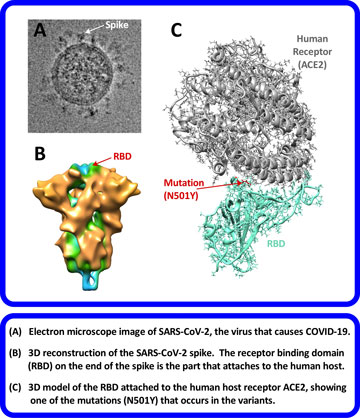
All this hard work paid off when initial results showed that it was possible for the virus mutation to be modelled and analyzed in 3D. Dr. Beniac looked specifically at the N501Y mutation found in the Alpha, Beta, Gamma, and Omicron variants and was able to detect an increase in binding capacity for this mutation.
An early warning device
This modelling system was developed with the intention of filling the gap in time between the sequencing of variants (which provides genetic information about the changes in the virus), and when epidemiological data (which is data on the population distribution of a disease by neighbourhood, city, province, country or globally) starts to become available. As new variants like Omicron continue to emerge, these receptor binding domain models are an additional tool scientists use to understand in part how the virus behaves, how it may be transmitted and how our bodies react to it. “This is meant to be a directional tool, like a compass, that helps us anticipate the effects of the virus when it has this mutation and sends us in a particular direction when it comes to public health policy,” says Dr. Beniac. “We can think of it as an early warning device.”
These models can help push the research of emerging variants in a certain direction and support decision-making related to public health measures. For example, if the model shows that a certain mutation found in a variant doesn’t affect binding at all, yet the variant is becoming dominant in a community, it can help shed insight as to why and narrow the cause for scientists—maybe it is instead an issue of immune evasion. In addition, mutations elsewhere in the variant can affect the spread of the virus, since this technique specifically tests for mutations in the receptor binding domain. Both variants of concern and immune evasion may affect the effectiveness of vaccines, so scientists are always on the lookout for it.
While this 3D model method is still being refined, Dr. Beniac hopes it will become a tool for future use. In the below figure, panel (C) shows an example of a model created using an atomic resolution structure.
“We may have more of these variants in the future,” says Dr. Beniac. “Any information we can get on these variants is essential to help guide us as new mutations arise.”
SARS-CoV-2 a Sense of Scale